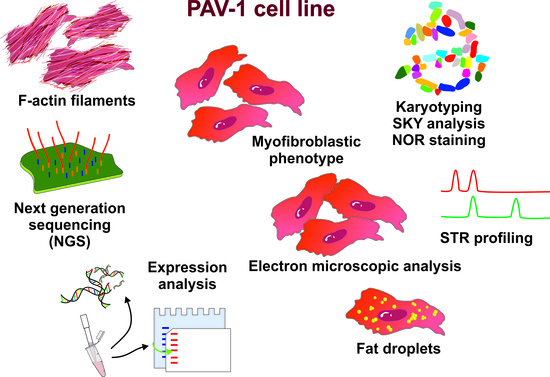
Genetic Characterization of Rat Hepatic Stellate Cell Line PAV-1
Abstract
:1. Introduction
2. Materials and Methods
2.1. Culturing of PAV-1 Cells
2.2. Electron Microscopy Analysis
2.3. Preparation of PAV-1 Metaphase Chromosomes and Karyotyping
2.4. In Situ Hybrdization, Spectral Imaging, and Nucleolar Organizer Region Staining
2.5. Short Tandem Repeat (STR) Profiling
2.6. Next Generation Sequencing and Data Analysis
2.7. Conventional Reverse Transcription Polymerase Chain Reaction
2.8. Western Blot Analysis
2.9. Rhodamine-Phalloidin Stain
2.10. Lipid Droplet Staining
3. Results
3.1. Phenotypic Characterization of PAV-1 Cells
3.2. Expression of Hepatic Stellate Cell Markers in PAV-1
3.2.1. Western Blot Analysis and Reverse Transcriptase PCR
3.2.2. Transcriptomic Analysis of PAV-1 Cells
3.3. Karyotype Analysis and Spectral Karyotyping of PAV-1 Cells
3.4. Spectral Karyotype Analysis
3.5. Short Tandem Repeat Profiling
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Acta2/α-SMA | α-smooth muscle actin |
| Aldh | aldehyde dehydrogenase |
| Bco | β-carotene oxygenase |
| CO1 | cytochrome c oxidase subunit 1 |
| Col | collagen |
| Crabp | cellular retinoic acid binding protein |
| DAPI | 4′,6-diamidino-2-phenylindole |
| DMEM | Dulbecco’s modified eagle medium |
| Fn1 | fibronectin |
| GAPDH | glyceraldehyde 3-phosphate dehydrogenase |
| GFAP | glial fibrillary acidic protein |
| GMO | genetically modified organism |
| HNF-4α | hepatocyte nuclear factor 4α |
| HRP | horseradish peroxidase |
| HSC(s) | hepatic stellate cell(s) |
| ICLAC | International Cell Line Authentication Committee |
| NIH | National Institutes of Health |
| NOR | nucleolus organizer regions |
| RAR | retinoid acid receptor |
| RBP | retinol binding protein |
| Rdh | retinol dehydrogenase |
| RT-PCR | conventional reverse transcription polymerase chain reaction |
| RT-qPCR | real-time quantitative PCR |
| RXR | retinoid X receptor |
| SKY | spectral karyotyping |
| SNP | single nucleotide polymorphism |
| STR | short tandem repeat |
| SV40T | Simian virus large T-antigen |
| TPM | transcripts per million |
| WGS | whole genome sequencing |
References
- Herrmann, J.; Gressner, A.M.; Weiskirchen, R. Immortal hepatic stellate cell lines: Useful tools to study hepatic stellate cell biology and function? J. Cell. Mol. Med. 2007, 11, 704–722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Friedman, S.L.; Weiskirchen, R. Working with Immortalized Hepatic Stellate Cell Lines. Methods Mol. Biol. 2023, 2669, 129–162. [Google Scholar] [CrossRef] [PubMed]
- Magness, S.T.; Bataller, R.; Yang, L.; Brenner, D. A dual reporter gene transgenic mouse demonstrates heterogeneity in hepatic fibrogenic cell populations. Hepatology 2004, 40, 1151–1159. [Google Scholar] [CrossRef]
- Krenkel, O.; Hundertmark, J.; Ritz, T.P.; Weiskirchen, R.; Tacke, F. Single Cell RNA Sequencing Identifies Subsets of Hepatic Stellate Cells and Myofibroblasts in Liver Fibrosis. Cells 2019, 8, 503. [Google Scholar] [CrossRef] [Green Version]
- Payen, V.L.; Lavergne, A.; Sarika, N.A.; Colonval, M.; Karim, L.; Deckers, M.; Najimi, M.; Coppieters, W.; Charloteaux, B.; Sokal, E.M.; et al. Single-cell RNA sequencing of human liver reveals hepatic stellate cell heterogeneity. JHEP Rep. 2021, 3, 100278. [Google Scholar] [CrossRef]
- Zhang, W.; Conway, S.J.; Liu, Y.; Snider, P.; Chen, H.; Gao, H.; Liu, Y.; Isidan, K.; Lopez, K.J.; Campana, G.; et al. Heterogeneity of Hepatic Stellate Cells in Fibrogenesis of the Liver: Insights from Single-Cell Transcriptomic Analysis in Liver Injury. Cells 2022, 10, 2129. [Google Scholar] [CrossRef]
- Yang, W.; He, H.; Wang, T.; Su, N.; Zhang, F.; Jiang, K.; Zhu, J.; Zhang, C.; Niu, K.; Wang, L.; et al. Single-Cell Transcriptomic Analysis Reveals a Hepatic Stellate Cell–Activation Roadmap and Myofibroblast Origin During Liver Fibrosis in Mice. Hepatology 2021, 74, 2774–2790. [Google Scholar] [CrossRef] [PubMed]
- Kimura, Y.; Koyama, Y.; Taura, K.; Kudoh, A.; Echizen, K.; Nakamura, D.; Li, X.; Nam, N.H.; Uemoto, Y.; Nishio, T.; et al. Characterization and role of collagen gene expressing hepatic cells following partial hepatectomy in mice. Hepatology 2022, 77, 443–455. [Google Scholar] [CrossRef] [PubMed]
- Russell, W.M.S.; Burch, R.L. The Principles of Humane Experimental Technique; Methuen: London, UK, 1959. [Google Scholar]
- Weiskirchen, R. Established Liver Cell Lines: Are You Sure to Have the Right Ones? Livers 2022, 2, 171–177. [Google Scholar] [CrossRef]
- Quevedo, R.; Smirnov, P.; Tkachuk, D.; Ho, C.; El-Hachem, N.; Safikhani, Z.; Pugh, T.J.; Haibe-Kains, B. Assessment of Genetic Drift in Large Pharmacogenomic Studies. Cell Syst. 2020, 11, 393–401.e2. [Google Scholar] [CrossRef] [PubMed]
- Horbach, S.P.J.M.; Halffman, W. The ghosts of HeLa: How cell line misidentification contaminates the scientific literature. PLoS ONE 2017, 12, e0186281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weill, F.-X.; Blazejewski, S.; Blanc, J.F.; Huet, S.; Gauthier, J.M.; Neaud, V.; Olaso, E.; Dubuisson, L.; Azais-Braesco, V.; Vidal-Vanaclocha, F.; et al. Characterization of a new human liver myofibroblast cell line: Transcriptional regulation of plasminogen activator inhibitor type I by transforming growth factor beta 1. Lab. Investig. 1997, 77, 63–70. [Google Scholar] [PubMed]
- MacLeod, R.A.; Dirks, W.G.; Matsuo, Y.; Kaufmann, M.; Milch, H.; Drexler, H.G. Widespread intraspecies cross-contamination of human tumor cell lines arising at source. Int. J. Cancer 1999, 83, 555–563. [Google Scholar] [CrossRef]
- The International Cell Line Authentication Committee. Available online: https://iclac.org/ (accessed on 17 November 2022).
- Souren, N.Y.; E Fusenig, N.; Heck, S.; Dirks, W.G.; Capes-Davis, A.; Bianchini, F.; Plass, C. Cell line authentication: A necessity for reproducible biomedical research. EMBO J. 2022, 41, e111307. [Google Scholar] [CrossRef]
- Almeida, J.L.; Cole, K.D.; Plant, A.L. Standards for Cell Line Authentication and Beyond. PLoS Biol. 2016, 14, e1002476. [Google Scholar] [CrossRef]
- Weiskirchen, R.; Weimer, J.; Meurer, S.K.; Kron, A.; Seipel, B.; Vater, I.; Arnold, N.; Siebert, R.; Xu, L.; Friedman, S.L.; et al. Genetic Characteristics of the Human Hepatic Stellate Cell Line LX-2. PLoS ONE 2013, 8, e75692. [Google Scholar] [CrossRef] [Green Version]
- Nanda, I.; Steinlein, C.; Haaf, T.; Buhl, E.M.; Grimm, D.G.; Friedman, S.L.; Meurer, S.K.; Schröder, S.K.; Weiskirchen, R. Genetic Characterization of Rat Hepatic Stellate Cell Line HSC-T6 for In Vitro Cell Line Authentication. Cells 2022, 11, 1783. [Google Scholar] [CrossRef]
- Nanda, I.; Schröder, S.K.; Steinlein, C.; Haaf, T.; Buhl, E.M.; Grimm, D.G.; Weiskirchen, R. Rat Hepatic Stellate Cell Line CFSC-2G: Genetic Markers and Short Tandem Repeat Profile Useful for Cell Line Authentication. Cells 2022, 11, 2900. [Google Scholar] [CrossRef]
- Schröder, S.K.; Schüler, H.M.; Petersen, K.V.; Tesauro, C.; Knudsen, B.R.; Pedersen, F.S.; Krus, F.; Buhl, E.M.; Roeb, E.; Roderfeld, M.; et al. Genetic and Molecular Characterization of the Immortalized Murine Hepatic Stellate Cell Line GRX. Cells 2022, 11, 1504. [Google Scholar] [CrossRef]
- Meurer, S.K.; Brenner, D.A.; Weiskirchen, R. Multiplex short tandem repeat profiling of immortalized hepatic stellate cell line Col-GFP HSC. PLoS ONE 2022, 17, e0274219. [Google Scholar] [CrossRef]
- Sauvant, P.; Sapin, V.; Abergel, A.; Schmidt, C.K.; Blanchon, L.; Alexandre-Gouabau, M.-C.; Rosenbaum, J.; Bommelaer, G.; Rock, E.; Dastugue, B.; et al. PAV-1, a new rat hepatic stellate cell line converts retinol into retinoic acid, a process altered by ethanol. Int. J. Biochem. Cell Biol. 2002, 34, 1017–1029. [Google Scholar] [CrossRef] [PubMed]
- Sauvant, P.; Abergel, A.; Partier, A.; Alexandre-Gouabau, M.-C.; Rock, E.; Sion, B.; Motta, C.; Sapin, V.; Azaïs-Bresco, V. Treatment of the rat hepatic stellate cell line, PAV-1, by retinol and palmitic acid leads to a convenient model to study retinoids metabolism. Biol. Cell 2002, 94, 401–408. [Google Scholar] [CrossRef] [PubMed]
- Sumner, A.T. A simple technique for demonstrating centromeric heterochromatin. Exp. Cell Res. 1972, 75, 304–306. [Google Scholar] [CrossRef] [PubMed]
- Hamta, A.; Adamovic, T.; Samuelson, E.; Helou, K.; Behboudi, A.; Levan, G. Chromosome ideograms of the laboratory rat (Rattus norvegicus) based on high-resolution banding, and anchoring of the cytogenetic map to the DNA sequence by FISH in sample chromosomes. Cytogenet. Genome Res. 2006, 115, 158–168. [Google Scholar] [CrossRef] [PubMed]
- Buwe, A.; Steinlein, C.; Koehler, M.; Bar-Am, I.; Katzin, N.; Schmid, M. Multicolor spectral karyotyping of rat chromosomes. Cytogenet. Genome Res. 2003, 103, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Goodpasture, C.; Bloom, S.E. Visualization of nucleolar organizer regions in mammalian chromosomes using silver staining. Chromosoma 1975, 53, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Almeida, J.L.; Hill, C.R.; Cole, K.D. Mouse cell line authentication. Cytotechnology 2014, 66, 133–147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Almeida, J.L.; Dakic, A.; Kindig, K.; Kone, M.; Letham, D.L.D.; Langdon, S.; Peat, R.; Holding-Pillai, J.; Hall, E.M.; Ladd, M.; et al. Interlaboratory study to validate a STR profiling method for intraspecies identification of mouse cell lines. PLoS ONE 2019, 14, e0218412. [Google Scholar] [CrossRef] [Green Version]
- The Cellosaurus STR Similarity Search Tool (CLASTR 1.4.4). Available online: https://web.expasy.org/cellosaurus-str-search/ (accessed on 17 November 2022).
- Bairoch, A. The Cellosaurus, a Cell-Line Knowledge Resource. J. Biomol. Tech. 2018, 29, 25–38. [Google Scholar] [CrossRef]
- Borkham-Kamphorst, E.; van de Leur, E.; Zimmermann, H.W.; Karlmark, K.R.; Tihaa, L.; Haas, U.; Tacke, F.; Berger, T.; Mak, T.W.; Weiskirchen, R. Protective effects of lipocalin-2 (LCN2) in acute liver injury suggest a novel function in liver homeostasis. Biochim. Biophys. Acta 2013, 1832, 660–673. [Google Scholar] [CrossRef] [Green Version]
- Schröder, S.K.; Asimakopoulou, A.; Tillmann, S.; Koschmieder, S.; Weiskirchen, R. TNF-α controls Lipocalin-2 expression in PC-3 prostate cancer cells. Cytokine 2020, 135, 155214. [Google Scholar] [CrossRef] [PubMed]
- Schröder, S.K.; Tag, C.G.; Weiskirchen, S.; Weiskirchen, R. Phalloidin Staining for F-Actin in Hepatic Stellate Cells. Methods Mol. Med. 2023, 2669, 55–66. [Google Scholar] [CrossRef]
- Drexler, H.G.; Uphoff, C.C. Mycoplasma contamination of cell cultures: Incidence, sources, effects, detection, elimination, prevention. Cytotechnology 2002, 39, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Pamies, D.; Leist, M.; Coecke, S.; Bowe, G.; Allen, D.G.; Gstraunthaler, G.; Bal-Price, A.; Pistollato, F.; de Vries, R.B.M.; Hogberg, H.T.; et al. Guidance document on Good Cell and Tissue Culture Practice 2.0 (GCCP 2.0). Altex 2022, 39, 30–70. [Google Scholar] [CrossRef]
- Blaner, W.S.; O’Byrne, S.M.; Wongsiriroj, N.; Kluwe, J.; D’Ambrosio, D.M.; Jiang, H.; Schwabe, R.F.; Hillman, E.M.; Piantedosi, R.; Libien, J. Hepatic stellate cell lipid droplets: A specialized lipid droplet for retinoid storage. Biochim. Biophys. Acta 2009, 1791, 467–473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Furr, H.C.; Clifford, A.J.; Smith, L.M.; Olson, J.A. The Effect of Dietary Fatty Acid Composition on Liver Retinyl Ester (Vitamin A Ester) Composition in the Rat. J. Nutr. 1989, 119, 581–585. [Google Scholar] [CrossRef]
- Abergel, A.; Sapin, V.; Dif, N.; Chassard, C.; Darcha, C.; Marcand-Sauvant, J.; Gaillard-Martinie, B.; Rock, E.; Dechelotte, P.; Sauvant, P. Growth Arrest and Decrease of α-SMA and Type I Collagen Expression by Palmitic Acid in the Rat Hepatic Stellate Cell Line PAV-1. Dig. Dis. Sci. 2006, 51, 986–995. [Google Scholar] [CrossRef]
- Walesky, C.; Apte, U. Role of Hepatocyte Nuclear Factor 4α (HNF4α) in Cell Proliferation and Cancer. Gene Expr. 2015, 16, 101–108. [Google Scholar] [CrossRef] [Green Version]
- Buniatian, G.; Hamprecht, B.; Gebhardt, R. Glial fibrillary acidic protein as a marker of perisinusoidal stellate cells that can dis-tinguish between the normal and myofibroblast-like phenotypes. Biol. Cell. 1996, 87, 65–73. [Google Scholar] [CrossRef]
- Acharya, P.; Chouhan, K.; Weiskirchen, S.; Weiskirchen, R. Cellular Mechanisms of Liver Fibrosis. Front. Pharmacol. 2021, 12, 671640. [Google Scholar] [CrossRef]
- Dollé, M.E.; Giese, H.; Hopkins, C.L.; Martus, H.J.; Hausdorff, J.M.; Vijg, J. Rapid accumulation of genome rearrangements in liver but not in brain of old mice. Nat. Genet. 1997, 17, 431–434. [Google Scholar] [CrossRef]
- Geraghty, R.J.; Capes-Davis, A.; Davis, J.M.; Downward, J.; Freshney, R.I.; Knezevic, I.; Lovell-Badge, R.; Masters, J.R.W.; Meredith, J.; Stacey, G.N.; et al. Guidelines for the use of cell lines in biomedical research. Br. J. Cancer 2014, 111, 1021–1046. [Google Scholar] [CrossRef] [Green Version]
- Vaught, J.; Korch, C.T. The Continuing Saga of Cell Line Misidentification. Biopreserv. Biobanking 2021, 19, 357–358. [Google Scholar] [CrossRef] [PubMed]
- Babic, Z.; Capes-Davis, A.; E Martone, M.; Bairoch, A.; Ozyurt, I.B.; Gillespie, T.H.; E Bandrowski, A. Incidences of problematic cell lines are lower in papers that use RRIDs to identify cell lines. Elife 2019, 8, e41676. [Google Scholar] [CrossRef] [PubMed]
- Schweppe, R.E.; Klopper, J.P.; Korch, C.; Pugazhenthi, U.; Benezra, M.; Knauf, J.A.; Fagin, J.A.; Marlow, L.A.; Copland, J.A.; Smallridge, R.C.; et al. Deoxyribonucleic Acid Profiling Analysis of 40 Human Thyroid Cancer Cell Lines Reveals Cross-Contamination Resulting in Cell Line Redundancy and Misidentification. J. Clin. Endocrinol. Metab. 2008, 93, 4331–4341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, Y.; Liu, Y.; Zheng, C.; Shen, C. Investigation of Cross-Contamination and Misidentification of 278 Widely Used Tumor Cell Lines. PLoS ONE 2017, 12, e0170384. [Google Scholar] [CrossRef] [Green Version]
- Mirabelli, P.; Coppola, L.; Salvatore, M. Cancer Cell Lines Are Useful Model Systems for Medical Research. Cancers 2019, 11, 1098. [Google Scholar] [CrossRef] [Green Version]
- Sauvant, P.; Sapin, V.; Partier, A.; Abergel, A.; Rosenbaum, J.; Azaïs-Braesco, V. Validation of a rat hepatic stellate cell line (PAV-1) to study vitamin A metabolism: Comparison with primary hepatic stellate cells. In Cells of the Hepatic Sinusoid, Proceedings of the 10th International Symposium on Cells of the Hepatic Sinusoid, Southampton, UK, 3–7 September 2000; Wisse, E., Knook, D.L., de Zanger, R., Arthur, M.J.P., Eds.; The Kupffer Cell Foundation: Leiden, The Netherlands, 2000; Volume 8, pp. 75–77. ISBN 90-803856-3-8. [Google Scholar]
- Sauvant, P.; Abergel, A.; Rosenbaum, J. Ethanol alters retinoic acid production in hepatic stellate cell line (PAV-1). Hepatology 2000, 32, 193A. [Google Scholar]
- Abergel, A.; Sauvant, P.; Sapin, V.; Dieu, H.; Ferrand, C.; Chassard, C.; Rock, E. Palmitic acid is an inhibitor of the proliferation of an hepatic stellate cell line, PAV-1. Hepatology 2001, 34, 521A. [Google Scholar]
- Chen, G.; Weiskirchen, S.; Weiskirchen, R. Vitamin A: Too good to be bad? Front. Pharmacol. 2023, 14, 1186336. [Google Scholar] [CrossRef]
- Feinstein, D.L.; Weinmaster, G.A.; Milner, R.J. Isolation of cDNA clones encoding rat glial fibrillary acidic protein: Expression in astrocytes and in Schwann cells. J. Neurosci. Res. 1992, 32, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Lim, M.C.C.; Maubach, G.; Lang, Z. Glial fibrillary acidic protein splice variants in hepatic stellate cells—Expression and regulation. Mol. Cells 2008, 25, 376–384. [Google Scholar] [PubMed]
- Xu, L.; Hui, A.Y.; Albanis, E.; Arthur, M.J.; O’byrne, S.M.; Blaner, W.S.; Mukherjee, P.; Friedman, S.L.; Eng, F.J. Human hepatic stellate cell lines, LX-1 and LX-2: New tools for analysis of hepatic fibrosis. Gut 2005, 54, 142–151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Horie, S.; Kitamura, Y.; Kawasaki, H.; Terada, T. Inhibitory effects of antisense oligonucleotides on the expression of procollagen type III gene in mouse hepatic stellate cells transformed by simian virus 40. Pathol. Int. 2000, 50, 937–944. [Google Scholar] [CrossRef]
- Kitamura, Y.; Tanigawa, T.; Katsumoto, T.; Tomita, K.; Wang, H.; Hirai, K.; Ichihara, K.; Terada, T. Cell growth and differentiation of a novel mouse Ito (fat-storing) cell line transformed by a temperature-sensitive mutant of simian virus 40. Hepatology 1997, 26, 323–329. [Google Scholar] [CrossRef]
- Miura, N.; Kanayama, Y.; Nagai, W.; Hasegawa, T.; Seko, Y.; Kaji, T.; Naganuma, A. Characterization of an immortalized hepatic stellate cell line established from metallothionein-null mice. J. Toxicol. Sci. 2006, 31, 391–398. [Google Scholar] [CrossRef] [Green Version]
- Vogel, S.; Piantedosi, R.; Frank, J.; Lalazar, A.; Rockey, D.C.; Friedman, S.L.; Blaner, W.S. An immortalized rat liver stellate cell line (HSC-T6): A new cell model for the study of retinoid metabolism in vitro. J. Lipid Res. 2000, 41, 882–893. [Google Scholar] [CrossRef]
- Meurer, S.K.; Alsamman, M.; Sahin, H.; Wasmuth, H.E.; Kisseleva, T.; Brenner, D.A.; Trautwein, C.; Weiskirchen, R.; Scholten, D. Overexpression of Endoglin Modulates TGF-β1-Signalling Pathways in a Novel Immortalized Mouse Hepatic Stellate Cell Line. PLoS ONE 2013, 8, e56116. [Google Scholar] [CrossRef] [Green Version]
- Guo, J.; Loke, J.; Zheng, F.; Hong, F.; Yea, S.; Fukata, M.; Tarocchi, M.; Abar, O.T.; Huang, H.; Sninsky, J.J.; et al. Functional linkage of cirrhosis-predictive single nucleotide polymorphisms of toll-like receptor 4 to hepatic stellate cell responses. Hepatology 2009, 49, 960–968. [Google Scholar] [CrossRef] [Green Version]
- Tsai, S.; Wear, D.J.; Shih, J.W.; Lo, S.C. Mycoplasmas and oncogenesis: Persistent infection and multistage malignant transformation. Proc. Natl. Acad. Sci. USA 1995, 92, 10197–10201. [Google Scholar] [CrossRef] [Green Version]
- Namiki, K.; Goodison, S.; Porvasnik, S.; Allan, R.W.; Iczkowski, K.A.; Urbanek, C.; Reyes, L.; Sakamoto, N.; Rosser, C.J. Persistent Exposure to Mycoplasma Induces Malignant Transformation of Human Prostate Cells. PLoS ONE 2009, 4, e6872. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, N.; Huang, X.; Jia, Y. Mycoplasma contamination affects cell characteristics and decreases the sensitivity of BV2 microglia to LPS stimulation. Cytotechnology 2019, 71, 623–634. [Google Scholar] [CrossRef] [PubMed]
- Romorini, L.; Riva, D.A.; Blüguermann, C.; Richardson, G.A.V.; Scassa, M.E.; Sevlever, G.E.; Miriuka, S.G. Effect of Antibiotics against Mycoplasma sp. on Human Embryonic Stem Cells Undifferentiated Status, Pluripotency, Cell Viability and Growth. PLoS ONE 2013, 8, e70267. [Google Scholar] [CrossRef] [PubMed]
- Weiskirchen, S.; Schröder, S.K.; Buhl, E.M.; Weiskirchen, R. A Beginner’s Guide to Cell Culture: Practical Advice for Preventing Needless Problems. Cells 2023, 12, 682. [Google Scholar] [CrossRef] [PubMed]









| Antibody | Cat. No. | Company | Expected Size (kDa) | Dilution | Clonality |
|---|---|---|---|---|---|
| α-SMA | CBL171-I | Sigma-Aldrich, Taufkirchen, Germany | 45 | 1:1000 | r mAb |
| β-actin | A5441 | Sigma-Aldrich | 43 | 1:10,000 | m mAb |
| Caveolin-1 | #3238 | Cell Signaling; Leiden, The Netherlands | 21–24 | 1:1000 | r pAb |
| Collagen I | NB600-408 | Novus Biologicals, Wiesbaden, Germany | 139 | 1:1000 | r mAb |
| Collagen IV | ab6586 | Abcam, Berlin, Germany | 161 | 1:1000 | r pAb |
| Desmin (D93F5) | #5332 | Cell Signaling | 53 | 1:1000 | r mAb |
| Fibronectin | AB1954 | Sigma-Aldrich | 262 | 1:3000 | r pAb |
| GAPDH (6C5) | sc-32233 | Santa Cruz Biotech., Santa Cruz, CA, USA | 37 | 1:1000 | m mAb |
| GFAP (2E1) | sc-33673 | Santa Cruz | 50 | 1:1000 | m mAb |
| GFAP (astrocyte marker) | ab7779 | Abcam | 50 (55) | 1:1000 | r pAb |
| HNF-4α (C-19) | sc-6556 | Santa Cruz | 54 (40) | 1:750 | p gAb |
| SV40T (v-300) | sc-20800 | Santa Cruz | 94 | 1:1000 | r pAb |
| Vimentin | ab92547 | Abcam | 54 | 1:3000 | r mAb |
| g anti-mIgG (H + L), HRP | #31430 | Invitrogen, ThermoFisher, Dreieich, Germany | NA | 1:5000 | g pAb |
| g anti-rIgG (H + L), HRP | #31460 | Invitrogen | NA | 1:5000 | g pAb |
| m anti-gIgG (H + L) HRP | #31400 | Invitrogen | NA | 1:5000 | m pAb |
| Transcript ID | Gene ID | Gene | Gene Description | PAV-1 (TPM) | HSC-T6 (TPM) | CFSC-2G (TPM) |
|---|---|---|---|---|---|---|
| ENSRNOT00000083468.1 | ENSRNOG00000058039.1 | Acta2 | actin alpha 2, smooth muscle | 58.2526 | 2.377 | 10.9245 |
| ENSRNOT00000005311.6 | ENSRNOG00000003897.6 | Col1a1 | collagen type I alpha 1 chain | 2470.3 | 2033.74 | 1408.42 |
| ENSRNOT00000004956.4 | ENSRNOG00000003357.4 | Col3a1 | collagen type III alpha 1 chain | 423.594 | 2128.9 | 64.4932 |
| ENSRNOT00000024430.5 | ENSRNOG00000018087.5 | Vim | vimentin | 7038.71 | 2054.81 | 3853.69 |
| ENSRNOT00000057585.4 | ENSRNOG00000014288.8 | Fn1 | fibronectin 1 | 1693.83 | 143.953 | 1383.81 |
| ENSRNOT00000019772.6 | 370.187 | 352.585 | 702.835 | |||
| ENSRNOT00000017486.7 | ENSRNOG00000012840.7 | Sparc | secreted protein acidic and cysteine rich | 2414.09 | 2188.34 | 1040.23 |
| ENSRNOT00000067011.2 | ENSRNOG00000003772.7 | Csrp2 | cysteine and glycine-rich protein 2 | 276.038 | 195.244 | 62.7395 |
| ENSRNOT00000080598.1 | 4.09374 | 4.42437 | 1.12115 | |||
| ENSRNOT00000013745.7 | ENSRNOG00000010208.7 | Timp1 | tissue inhibitor of metallopeptidase 1 | 646.539 | 5518.57 | 714.162 |
| ENSRNOT00000010180.5 | ENSRNOG00000007650.5 | Cd63 | Cd63 molecule | 2524.99 | 1639.48 | 2732.07 |
| ENSRNOT00000090381.1 | 85.4183 | 60.6936 | 106.408 | |||
| ENSRNOT00000011208.7 | ENSRNOG00000008301.7 | Tagln2 | transgelin 2 | 1999.07 | 754.93 | 642.605 |
| ENSRNOT00000034401.5 | ENSRNOG00000002919.9 | Gfap | glial fibrillary acidic protein | 0.0115619 | 0.60103 | 0.108417 |
| ENSRNOT00000093266.1 | 0 | 0 | 0 | |||
| ENSRNOT00000093167.1 | 0 | 0 | 0 | |||
| ENSRNOT00000042459.4 | ENSRNOG00000034254.4 | Actb | actin, beta | 3767.55 | 4379.5 | 3869.05 |
| ENSRNOT00000080216.1 | 1524.6 | 1820.7 | 1503.75 | |||
| ENSRNOT00000050443.4 | ENSRNOG00000018630.7 | LOC108351137 | glyceraldehyde-3-phosphate dehydrogenase | 2100.85 | 4598.86 | 3223.93 |
| ENSRNOT00000041328.3 | ENSRNOG00000030963.3 | LOC108351137 | glyceraldehyde-3-phosphate dehydrogenase | 1893.33 | 4243.2 | 2962.7 |
| Transcript ID | Gene ID | Gene | Gene Description | PAV-1 (TPM) | HSC-T6 (TPM) | CFSC-2G (TPM) |
|---|---|---|---|---|---|---|
| ENSRNOT00000008659.4 | ENSRNOG00000009972.7 | RARα | retinoic acid receptor, alpha | 34.5437 | 24.8697 | 26.5339 |
| ENSRNOT00000084644.1 | 25.8309 | 23.8544 | 20.4423 | |||
| ENSRNOT00000033048.6 | ENSRNOG00000024061.7 | RARβ | retinoic acid receptor, beta | 0.102317 | 0.541197 | 0.00872314 |
| ENSRNOT00000016801.5 | ENSRNOG00000012499.7 | RARγ | retinoic acid receptor, gamma | 67.7163 | 64.5919 | 49.0507 |
| ENSRNOT00000017096.7 | 17.0844 | 39.1824 | 25.6328 | |||
| ENSRNOT00000012892.4 | ENSRNOG00000009446.4 | RXRα | retinoid X receptor alpha | 75.5966 | 18.9052 | 41.2102 |
| ENSRNOT00000041613.5 | ENSRNOG00000000464.7 | RXRβ | retinoid X receptor beta | 9.88301 | 20.0384 | 15.9738 |
| ENSRNOT00000087670.1 | 1.0449 | 1.79029 | 1.53037 | |||
| ENSRNOT00000086978.1 | 0.43314 | 1.08069 | 1.02833 | |||
| ENSRNOT00000081588.1 | 2.00124 | 0.804912 | 1.61434 | |||
| ENSRNOT00000091182.1 | 1.43022 | 0.723898 | 0.347194 | |||
| ENSRNOT00000087895.1 | 0.824919 | 0.688452 | 0.0953106 | |||
| ENSRNOT00000084638.1 | 1.03353 | 0.499241 | 0.615142 | |||
| ENSRNOT00000079967.1 | 0.197368 | 0 | 0 | |||
| ENSRNOT00000077227.1 | ENSRNOG00000004537.6 | RXRγ | retinoid X receptor gamma | 0 | 0 | 0 |
| ENSRNOT00000006117.5 | 0 | 0 | 0.271271 | |||
| ENSRNOT00000018622.4 | ENSRNOG00000013794.4 | Rbp1 (CRBP1) | retinol binding protein 1 | 1.28709 | 25.3842 | 0 |
| ENSRNOT00000018755.6 | ENSRNOG00000013932.6 | Rbp2 (CRBP2) | retinol binding protein 2 | 0.695375 | 0.309974 | 1.64899 |
| ENSRNOT00000081756.1 | ENSRNOG00000053850.1 | Rdh5 (Rdh1) | retinol dehydrogenase 5 (formely 1) | 0.65424 | 0.0383589 | 0.693011 |
| ENSRNOT00000024000.6 | ENSRNOG00000017619.7 | ALDH1A1 | retinal dehydrogenase 1 | 0.00983515 | 0.291398 | 0.0251453 |
| ENSRNOT00000079115.1 | ENSRNOG00000055049.1 | ALDH1A2 | retinal dehydrogenase 2 | 0.0287604 | 15.2533 | 0 |
| ENSRNOT00000003182.5 | ENSRNOG00000002331.5 | ALDH3A1 | aldehyde dehydrogenase 3 (Tumor ALDH) | 240.833 | 21.5281 | 90.6901 |
| ENSRNOT00000001816.6 | ENSRNOG00000001344.7 | ALDH2 | mitochondrial aldehyde dehydrogenase | 522.001 | 528.489 | 76.4596 |
| ENSRNOT00000077461.1 | ENSRNOG00000002342.6 | ALDH3A2 | microsomal ALDH | 18.5197 | 8.59967 | 9.78843 |
| ENSRNOT00000066109.3 | 73.9895 | 37.999 | 42.67 |
| Passage No. | <Diploid | Around Diploid | Around Tetraploid |
|---|---|---|---|
| 5th | 1 | 83 | 17 |
| 10th | 0 | 54 | 46 |
| 15th | 1 | 39 | 61 |
| 20th | 1 | 31 | 69 |
| 30th | 2 | 39 | 59 |
| Cell | Chromosome Numbers | der(1)t(1;3) | der(1)t(1;18) | der(2)t(2;11p) | der(3)t(3;12) | der(4)t(4;6) | der(4)t(4;11p) | der(5)t(5;5) | der(6)t(6;11p) | der(6)t(6;12) | der(7)t(7;16) | der(7)t(7;17) | der(7)t(7;19) | der(10)t(10;13/14) | der(10)t(10;19) | der(11)t(11;17) | der(11)t(11;18) | der(11)t(11;20) | der(12)t(12;16) | der(12)t(12;18) | der(12)t(12;19) | der(12)t(12;20) | der(16)t(16;17) | der(16)t(16;19) | der(17)t(17;18) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 59,X | + | + | ++ | ++ | ||||||||||||||||||||
| 2 | 80,XXYY | + | ++ | ++ | + | + | ++ | + | |||||||||||||||||
| 3 | 77,XYYY | + | + | ++ | ++ | ++ | + | ||||||||||||||||||
| 4 | 39,XY | + | + | + | + | ||||||||||||||||||||
| 5 | 41,XY | ++ | + | + | + | ||||||||||||||||||||
| 6 | 77,XYY | + | ++ | ++ | ++ | + | + | ||||||||||||||||||
| 7 | 83,XXYY | + | ++ | ++ | + | ++ | ++ | ||||||||||||||||||
| 8 | 86,XXYY | + | ++ | ++ | ++ | + | + | ++ | + | + | + | ||||||||||||||
| 9 | 80,XXYY | + | ++ | ++ | + | + | ++ | + | |||||||||||||||||
| 10 | 69,XXYY | + | ++ | + | + | ++ | ++ | + | |||||||||||||||||
| 11 | 83,XXYY | + | ++ | ++ | + | ++ | ++ | ||||||||||||||||||
| 12 | 43,XY | + | + | + | + | ||||||||||||||||||||
| 13 | 81,XXYY | + | ++ | ++ | + | ++ | ++ | ||||||||||||||||||
| 14 | 43,XY | + | + | + | + | ||||||||||||||||||||
| 15 | 80,XXYY | + | ++ | + | ++ | ++ | |||||||||||||||||||
| 16 | 83,XXYY | + | ++ | ++ | + | ++ | ++ | + | |||||||||||||||||
| 17 | 81,XXYY | ++ | ++ | ++ | + | + | ++ | ++ | + | ||||||||||||||||
| 18 | 84,XXY | + | ++ | ++ | ++ | + | ++ | + | |||||||||||||||||
| 19 | 41,XY | + | + | + | + | + | + | ||||||||||||||||||
| 20 | 79,XXYYY | + | + | ++ | + | + | ++ |
| SN | Marker Name | Chromosomal Location | Allele Sizes (bp) in PAV-1 | Allele Sizes (bp) in CFSC-2G | Allele Sizes (bp) in HSC-T6 |
|---|---|---|---|---|---|
| 1 | 73 | 1 | 194 | 194, 203 | 194 |
| 2 | 8 | 2 | 234, 238 | 236 | 234 |
| 3 | 2 | 2 | 128 | 126 | 127 |
| 4 | 4 | 3 | 236, 238 | 268, 270 | 238 |
| 5 | 3 | 3 | 162 | 160, 182 | 160, 162 |
| 6 | 26 | 4 | 154 | 150 | 166 |
| 7 | 19 | 4 | 179 | 180 | 175 |
| 8 | 81 | 5 | 130 | 130, 134 | 130, 132 |
| 9 | 34 | 6 | 182, 187 | 184, 189 | 188 |
| 10 | 30 | 7 | 192 | 188, 192 | 186, 192 |
| 11 | 24 | 8 | 254, 259 | 260 | 249, 253 |
| 12 | 59 | 9 | 145, 148 | 145 | 143, 146, 180 |
| 13 | 62 | 9 | 166 | 166 | 177 |
| 14 | 1 | 10 | 96, 105 | 105 | 96 |
| 15 | 55 | 10 | 210, 218 | 210, 214 | 210, 218 |
| 16 | 36 | 11 | 222 | 222 | 234 |
| 17 | 67 | 11 | 165 | 154, 156 | 165 |
| 18 | 13 | 12 | 121 | 121 | 121, 135 |
| 19 | 35 | 13 | 203 | 197 | 197, 203 |
| 20 | 42 | 13 | 144, 156 | 125 | 127 |
| 21 | 70 | 14 | 158, 175 | 158, 175 | 175, 179 |
| 22 | 61 | 15 | 128 | 128 | 128 |
| 23 | 79 | 15 | 172 | 172, 180 | 172 |
| 24 | 90 | 16 | 175 | 159, 161 | 174 |
| 25 | 69 | 16 | 136, 139 | 138 | 139 |
| 26 | 78 | 17 | 147, 149 | 136, 151 | 147, 151 |
| 27 | 15 | 18 | 232 | 232 | 232 |
| 28 | 16 | 18 | 251 | 251, 260 | 247, 251 |
| 29 | 75 | 19 | 144, 184 | 144 | 144, 184 |
| 30 | 96 | 20 | 210 | 210 | 210, 212 |
| 31 | 91 | 20 | 211, 225 | 221 | 205, 211 |
| Characteristic | HSC-T6 | CFSC-2G | PAV-1 |
|---|---|---|---|
| Origin | Sprague Dawley | Wistar | Wistar |
| Sex | female 2 | male | male |
| Immortalization | SV40T | cirrhotic liver (CCl4) | spontaneous |
| Morphology | fibroblastic (spindle shaped) | fibroblastic (flat) | fibroblastic (spindle shaped) |
| Chromosome number | <43> | <62> | <43> |
| Derivative chromosomes | der (1), der (4), der (7) | der(1), der(2), der(4), der(6), der(7) der(10), der(14), der(17), der(19) | der (4), der (5), der (11), der (12) |
| Additional chromosomes | +4, +7 | +1, +2, +3, +4, +5, +6, +7, +8, +10 +13, +14, +17, +19 | +7, +11, +mar |
| Missing chromosomes | none | −15, −16 | none |
| STR profile | available [19] | available [20] | this study |
| Common HSC markers | see Table 2 | see Table 2 | see Table 2 |
| SV40T expression | positive | negative | negative |
| Special medium requirements | none | non-essential amino acids | none |
| Proposed field of study | general HSC aspects retinoid metabolism | general HSC aspects collagen expression and biology | fat metabolism retinol metabolism |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gäberlein, K.; Schröder, S.K.; Nanda, I.; Steinlein, C.; Haaf, T.; Buhl, E.M.; Sauvant, P.; Sapin, V.; Abergel, A.; Weiskirchen, R. Genetic Characterization of Rat Hepatic Stellate Cell Line PAV-1. Cells 2023, 12, 1603. https://0-doi-org.brum.beds.ac.uk/10.3390/cells12121603
Gäberlein K, Schröder SK, Nanda I, Steinlein C, Haaf T, Buhl EM, Sauvant P, Sapin V, Abergel A, Weiskirchen R. Genetic Characterization of Rat Hepatic Stellate Cell Line PAV-1. Cells. 2023; 12(12):1603. https://0-doi-org.brum.beds.ac.uk/10.3390/cells12121603
Chicago/Turabian StyleGäberlein, Kiara, Sarah K. Schröder, Indrajit Nanda, Claus Steinlein, Thomas Haaf, Eva M. Buhl, Patrick Sauvant, Vincent Sapin, Armand Abergel, and Ralf Weiskirchen. 2023. "Genetic Characterization of Rat Hepatic Stellate Cell Line PAV-1" Cells 12, no. 12: 1603. https://0-doi-org.brum.beds.ac.uk/10.3390/cells12121603